-
近年来,许多有机污染物在河流、地下水和湖泊中被发现,包括抗生素、农药和微塑料,并且含量维持在痕量浓度[1]。其中,抗生素和农药被广泛用于医疗和农业领域,这些污染物在痕量浓度下长期存在于水体中,会对饮用水造成严重危害[2]。长期过度使用抗生素导致细菌产生耐药性,从而降低抗生素对人类疾病的治疗效果[3]。因此,减少诱发抗性基因(antibiotic resistance genes, ARGs)的风险尤为重要。有研究[4]表明,氮循环细菌暴露在β-内酰胺类抗生素和寡营养环境下,成为主要抗性基因宿主。非抗生素化学物质,例如季铵化合物、微塑料可以诱导微生物应激反应,促进ARGs水平转移[5-6]。KURENBACH等[7]研究发现,除草剂可将抗生素的最小抑菌浓度提高6倍,促进微生物抗性基因产生。
活性炭具有发达的孔隙结构和不错的经济效益,被普遍用作饮用水高级处理的吸附剂。同时,活性炭可以与水体中微生物结合,利用微生物去除污染物,因此,活性炭通常被称为生物活性炭[8-9]。大量微生物附着于生物活性炭表面,ARGs的水平转移在微生物群落之间增强,从而引起耐药性细菌的增殖[10-11]。此外,有研究[12-13]表明,生物膜促进消毒过程中消毒副产物的产生,尤其是含氮类消毒副产物(nitrogenous disinfection byproducts formation potential, N-DBPs),究其原因是胞外聚合物(extracellular polymeric substance, EPS)、可溶性微生物产物和死细胞可以作为消毒副产物前驱体参与了反应。LI等[14]研究表明,环境胁迫促使微生物过度分泌EPS确保生存。蛋白质、多糖和核酸是EPS的主要成分,此外,蛋白质比多糖更容易促进N-DBPs的产生,因为其含有高含量的有机氮[15-16]。
LYU等[17-18]研究发现,吸附在双反应中心(dual-reaction-center, DRC)贫电子区的有机物能被高效降解,同时吸附在富电子区的H2O2被还原成羟基自由基或活性氧。有研究表明,微生物可以通过生物膜吸收不溶性电子供体提供的电子,例如,硫氧化硫化杆菌(Sulfobacillus thermosulfidooxidans)通过外膜蛋白从石墨表面电子供体吸收电子[19]。因此,电子从吸附在催化剂贫电子区的有机物到富电子区的微生物。这种滤料-微生物协同作用避免微生物从污染物中获得能量,削弱污染物对微生物的胁迫效应。
本研究通过浸渍和表面络合法制备了DRC催化剂(HCLL-S8-M),将其用于饮用水深度处理工艺。使用溶解性有机碳(dissolved organic carbon, DOC)的质量浓度、SUVA (specific ultraviolet absorbance)、高效凝胶排阻色谱(high-performance size exclusion chromatography, HPSEC)和三维荧光(excitation-emission matrix fluorescence, EEM)参数反映HCLL-S8-M滤柱对水质处理效果。通过实时荧光定量PCR (polymerase chain reaction)检测了微生物ARGs的相对丰度,使用气相色谱分析水体中N-DBPs生成势(N-DBPs formation potential, N-DBPs FP)。通过宏基因组学分析结合EPS的理化性质探讨微生物群落结构的变化,阐明催化剂、微生物之间的相互作用。探讨了在饮用水处理过程中,微生物协同HCLL-S8-M削弱复合微污染物对微生物胁迫效应的机制,为降低N-DBPs和ARGs风险提供理论依据。
-
根据已报道的方法合成氮掺杂类石墨烯络合Cu-Al2O3陶粒(HCLL-S8-M)催化剂[20],将0.58 g Al(NO3)3·9H2O和0.55 g Cu(NO3)2·3H2O均匀地分散3 mL水中。在水溶液中分散一定量的双氰胺,搅拌10 min后,将混合物倒入装有 10 g商用球形Al2O3颗粒(2 mm)的烧杯中。随后,向烧杯中加入NaOH使溶液呈碱性,并在100 ℃下干燥12 h,得到的固体样品在500 ℃下煅烧1 h。将市售的椰壳活性炭在10%的NaOH中浸泡2 h,然后用超纯水清洗至pH为中性。用10% HNO3进行上述同样的处理。最后,将椰壳活性炭在150 ℃的鼓风干燥箱中干燥24 h。
-
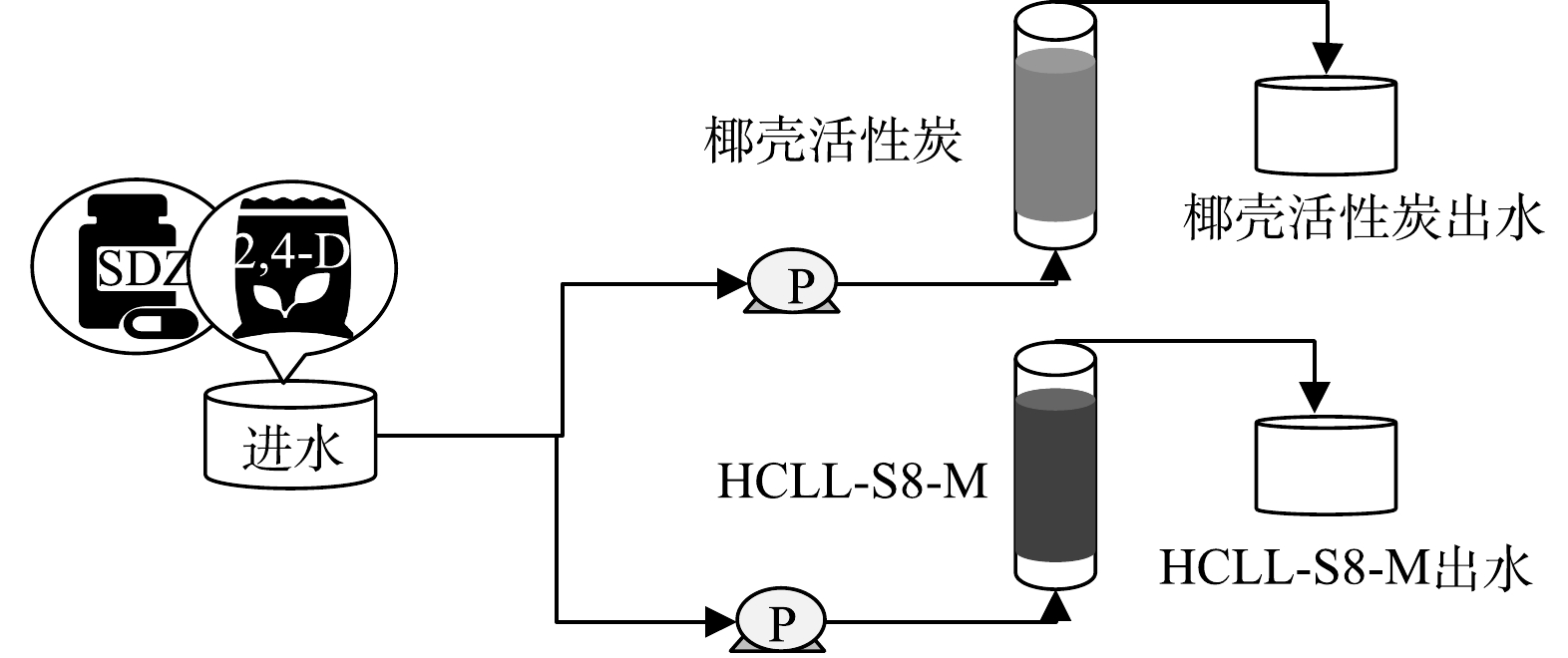
分别以HCLL-S8-M和椰壳活性炭为填充滤料制作2个反应器,模拟饮用水深度处理过程,装置示意图如图1所示。进水采用中国南方某饮用水处理厂经过砂滤处理后的出水。并且在进水中同时加入微污染物磺胺嘧啶(sulfadiazine, SDZ)和2,4-二氯苯氧乙酸(2,4-Dichlorophenoxyacetic acid, 2,4-D)各1 μg·L−1。进水从底部泵入反应器,空床停留时间为20 min,运行周期为46 d。收集不同滤料条件下的出水样品,进行化学和微生物分析。
-
DOC的质量浓度由总有机碳分析仪(TOC-L ASI-L,岛津,日本)测定;使用分光光度计(DR6000,哈希,美国)测量在254 nm波长下吸光度(UV254);特定波长的紫外吸光度(SUVA)计算公式为UV254/DOC×100[21];荧光分光光度计(F-7000,日立,日本)和液相色谱(1260 Infinity II,安捷伦,美国)分别测定EEM和HPSEC。N-DBPs FP根据已报道方法进行检测[21]。
-
EPS根据已报道方法进行提取,蛋白质和多糖质量浓度分别采用改良Lowry法和苯酚-硫酸法测定[21]。Zeta电位使用Zetasizer Nano(ZS90,马尔文,英国)测量。傅立叶变换红外光谱使用溴化钾粉末(Nicolet iS 10,赛默飞,美国)进行采集。
-
16S rRNA和ARGs通过实时荧光定量PCR(qPCR)检测。宏基因组学分析使用 Illumina NovaSeq/Hiseq Xten测序平台(因美纳,美国)。详细的qPCR流程、引物信息根据已报道方法进行检测[21]。BioProject: PRJNA1078285。
-
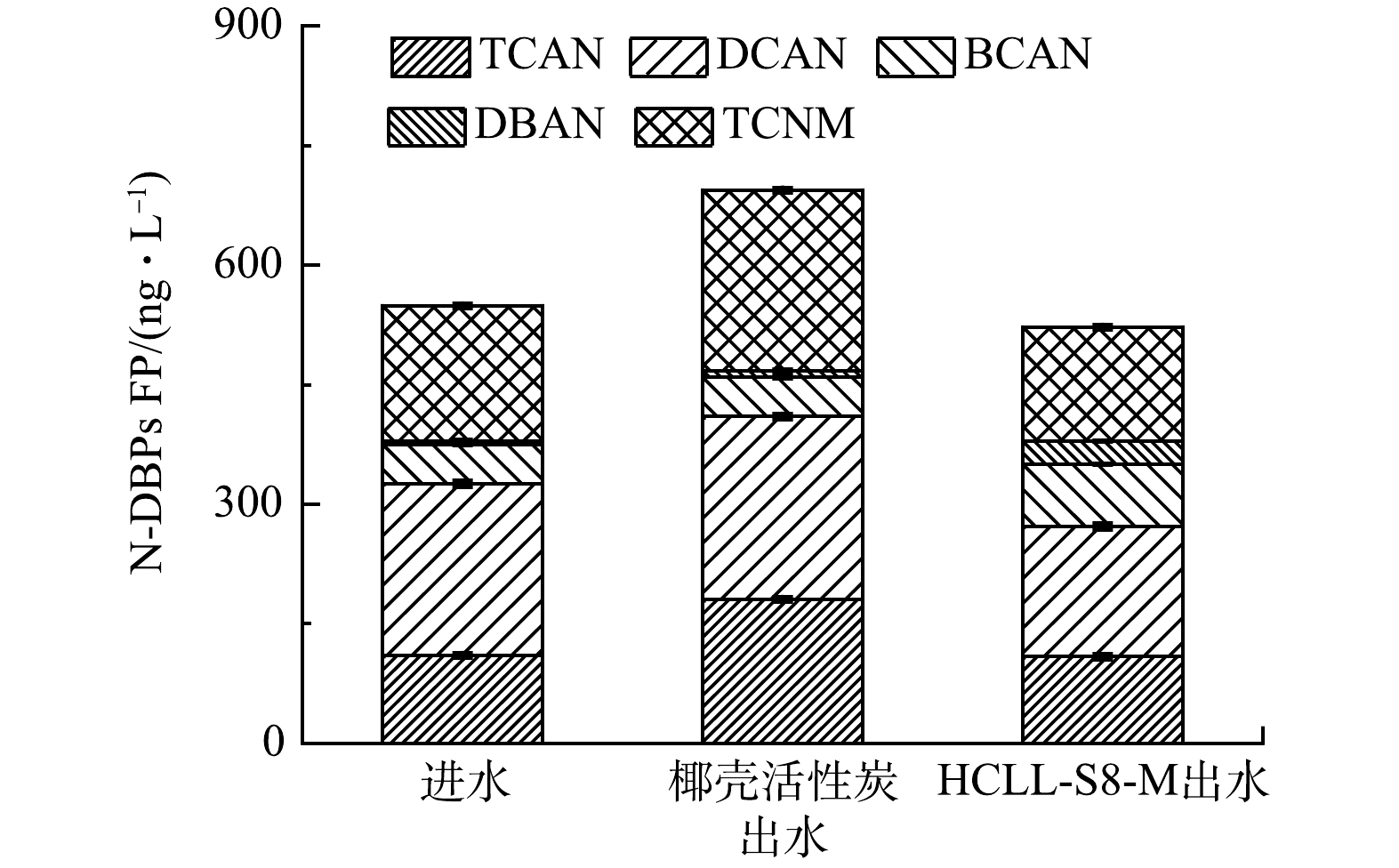
本研究检测不同滤料出水中卤代乙腈和卤代硝基甲烷的总生成势。其中包括三氯乙腈(trichloroacetonitrile, TCAN)、二氯乙腈(dichloroacetonitrile, DCAN)、溴氯乙腈(bromochloroacetonitrile, BCAN)、二溴乙腈(dibromoacetonitrile, DBAN)和三氯硝基甲烷(trichloronitromethane, TCNM)。图2表示N-DBPs FP在进水和出水的样品中质量浓度情况。
生物滤池运行46 d,HCLL-S8-M滤柱出水中N-DBPs FP质量浓度是522.30 ng·L−1,比椰壳活性炭滤柱出水中N-DBPs FP质量浓度降低171.70 ng·L−1,同时相比进水中N-DBPs FP质量浓度也下降26.44 ng·L−1。此外,椰壳活性炭滤柱出水比进水中N-DBPs FP增加145.30 ng·L−1。DCAN FP在所有样品中的质量浓度是最高的,这与文献报道的结果一致[22]。此外,在椰壳活性炭滤柱出水中,TCAN FP和TCNM FP质量浓度比HCLL-S8-M滤柱出水中(108.70 ng·L−1和143.40 ng·L−1)分别增加了71.50 ng·L−1和83.38 ng·L−1。结果表明,在复合微污染物条件下,HCLL-S8-M滤柱去除水体中N-DBPs前驱体的能力远高于椰壳活性炭滤柱。
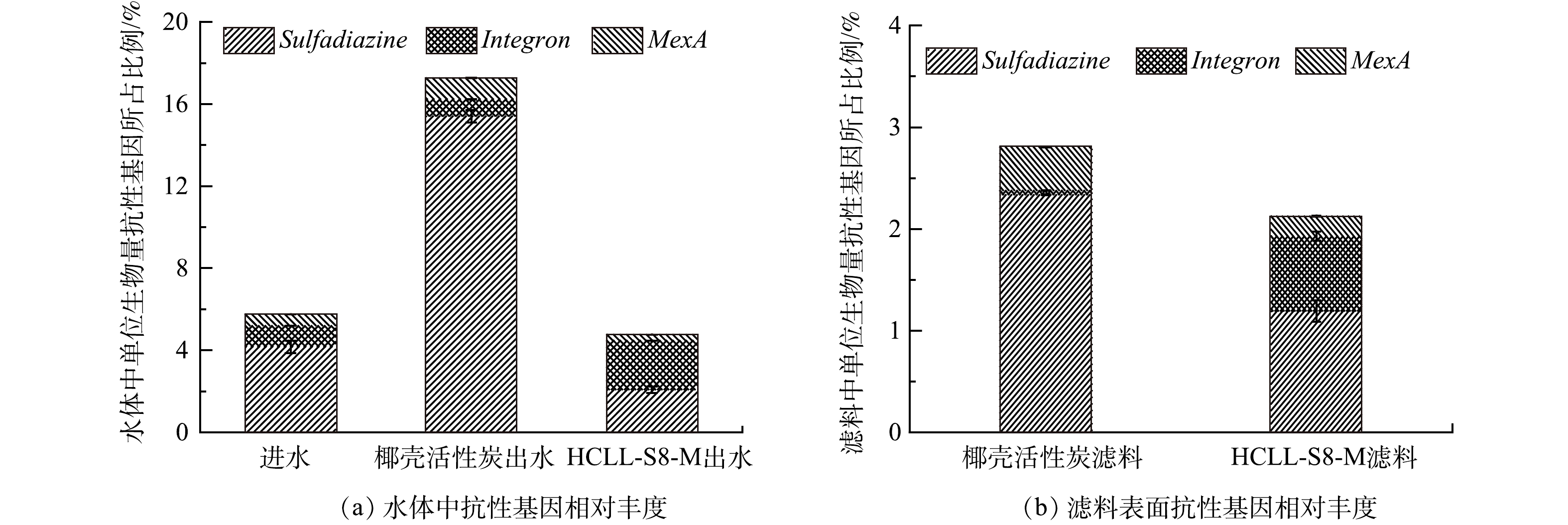
ARGs/16S rRNA比值表示ARGs的相对丰度,可以代表ARGs在样品微生物种群中的相对含量。图3表明,椰壳活性炭滤柱出水中ARGs的相对丰度是17.27%,比HCLL-S8-M滤柱出水(4.77%)和进水(5.75%)中ARGs的相对丰度分别高约4倍和3倍。此外,2种滤料表面ARGs的相对丰度也呈现相同趋势。在HCLL-S8-M滤柱中磺胺类ARGs的相对丰度低于椰壳活性炭滤柱,MexA的相对丰度比椰壳活性炭滤柱低2倍,外排泵可以主动将包括抗生素在内的多种底物排出胞外,促进抗性基因的产生[23]。此外,有研究[24-25]表明,生物膜不仅可以作为N-DBPs的前驱体,还能促进水中微生物之间的水平基因转移,导致ARGs的相对丰度增加。因此,HCLL-S8-M滤柱可以有效降低复合微污染物对微生物的胁迫效应。
-
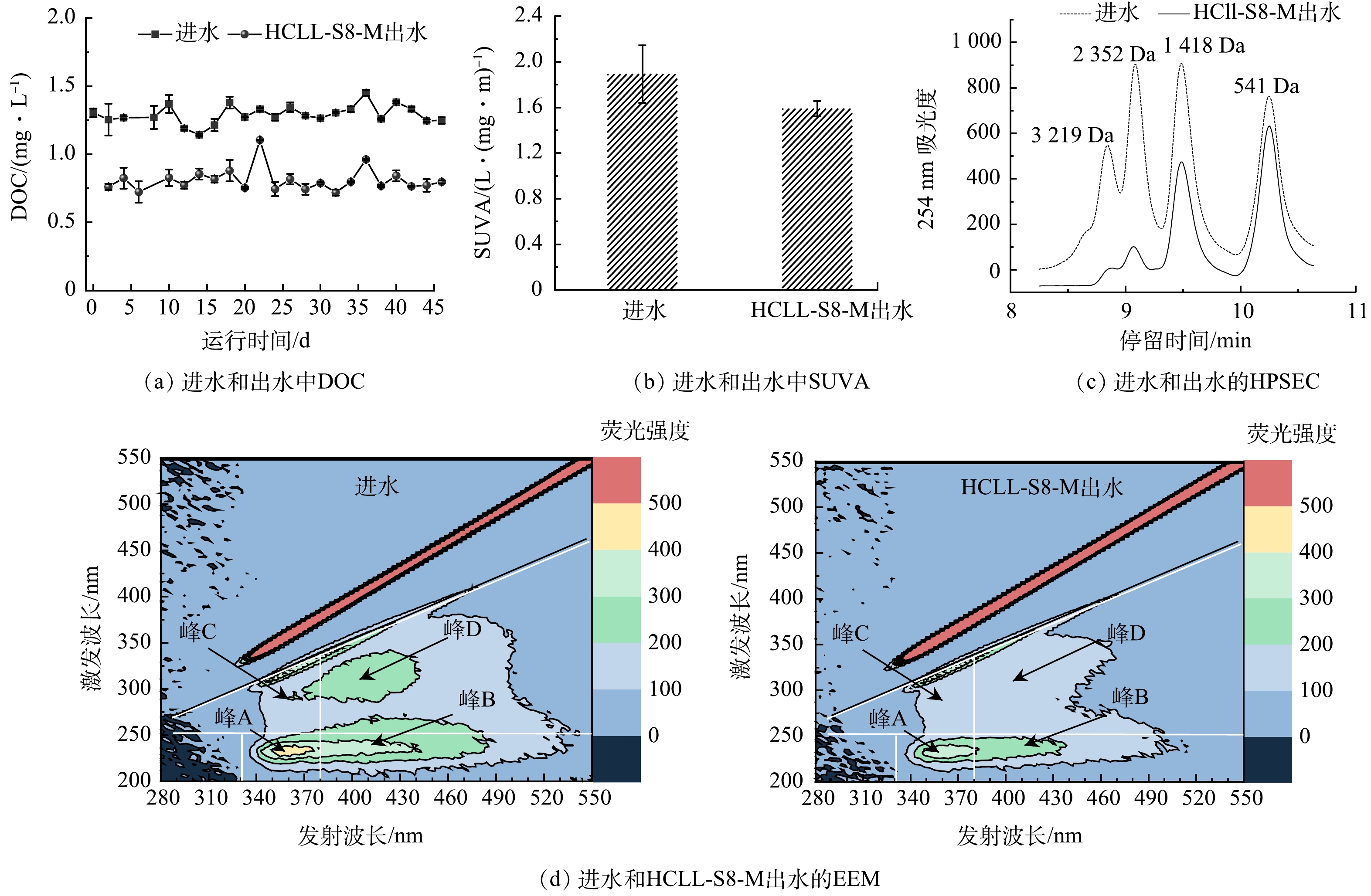
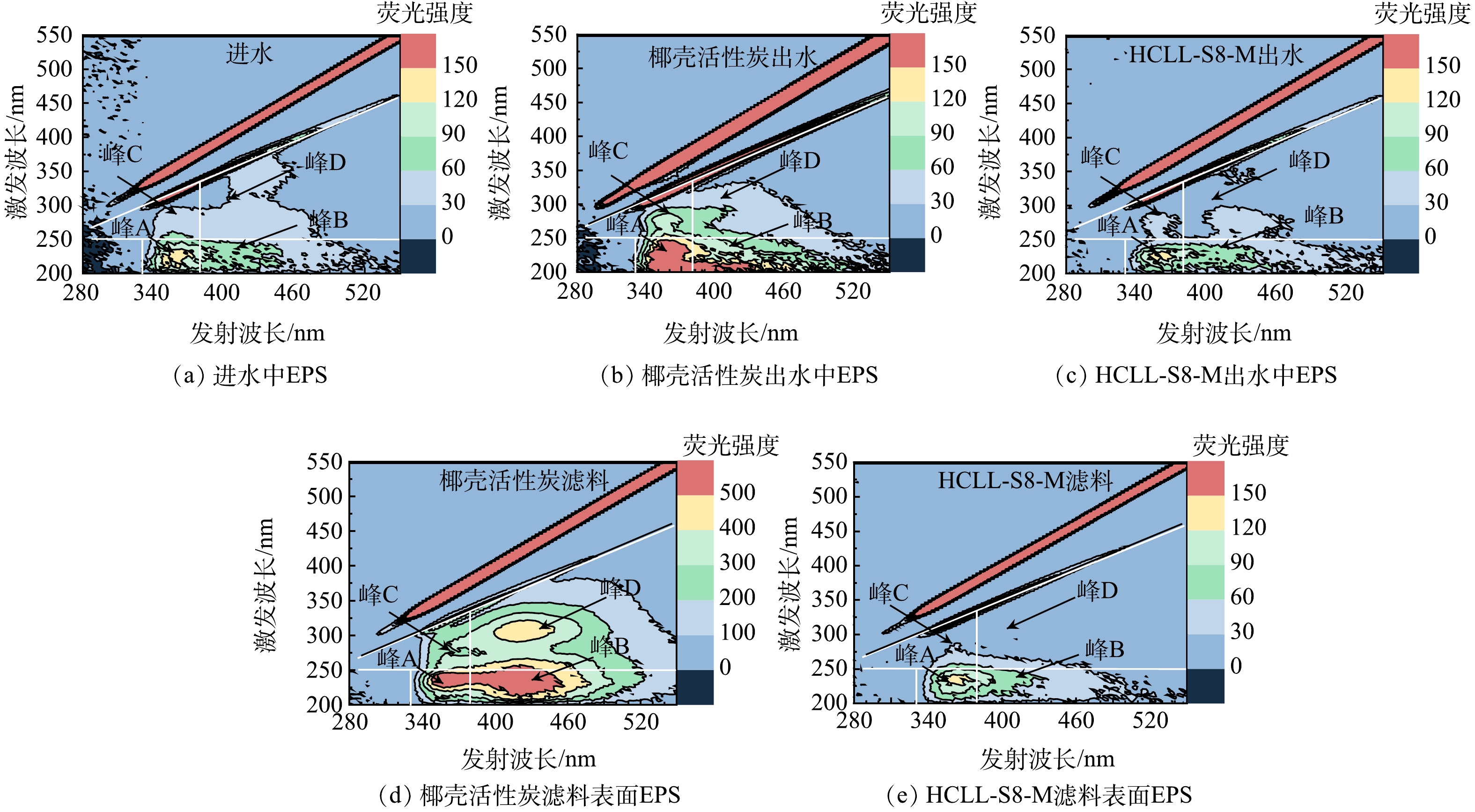
本实验连续运行46 d,由图4可见,经过HCLL-S8-M滤柱处理后,DOC质量浓度由1.14~1.45 mg·L−1下降到大约0.79 mg·L−1。出水的SUVA(1.59 L·(mg·m)−1)低于进水(1.89 L·(mg·m)−1),HCLL-S8-M滤柱可以有效去除DBPs前驱体[26]。根据HPSEC色谱图,观察到HCLL-S8-M滤柱处理后DOC中的高分子质量组份明显下降(保留时间小于10.0 min > 1 200 Da)。基于对EEM进行积分通常得到5个区域为:芳香蛋白质类物质(峰A:Ex/Em=220~250 nm/280~380 nm)、富里酸类物质(峰B:Ex/Em=200~250 nm/380~480 nm)、可溶性微生物产物类物质(峰C:Ex/Em=250~340 nm/280~380 nm)和腐殖酸类物质(峰D:Ex/Em=250~370 nm/380~480 nm)[27]。明显可见,经过HCLL-S8-M滤柱过滤后,所有区域的荧光强度均下降(图4(d)),这与DOC质量浓度的变化结果一致(图4(a))。结果表明,HCLL-S8-M滤柱对饮用水源广泛存在的DOC有很好的去除效果,同时具有胁迫作用的有机微污染物的随之去除。进一步说明HCLL-S8-M滤柱可以有效降低复合微污染物对微生物的胁迫效应,从而降低消毒副产物的生成量和抗性基因风险。
-
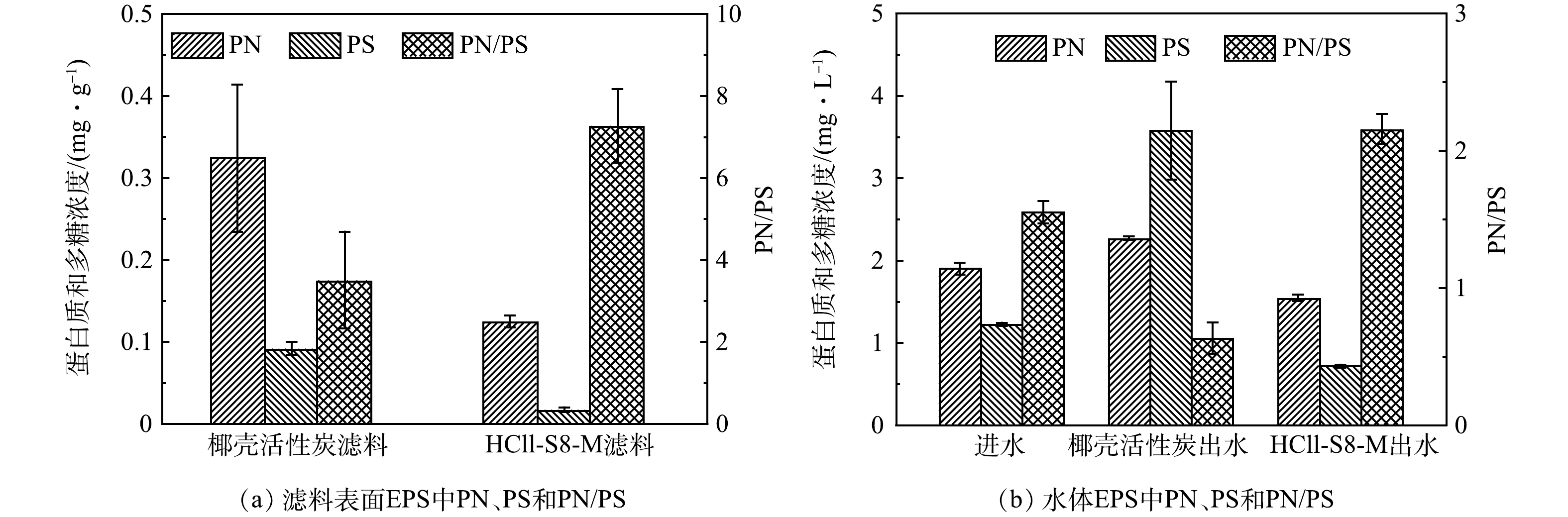
由图5可见,HCLL-S8-M滤料表面蛋白质含量比椰壳活性炭滤料表面蛋白质低0.20 mg·g−1,HCLL-S8-M滤柱出水EPS中蛋白质质量浓度(1.55 mg·L−1)也低于椰壳活性炭滤柱出水中蛋白质质量浓度(2.27 mg·L−1)。由图6可见,无论是在水体中还是滤料表面,HCLL-S8-M滤柱中EPS的芳香蛋白质类物质强度远低于椰壳活性炭滤柱,表明蛋白质可能在DBPs和ARGs的风险中发挥了重要作用。丰富的蛋白质类物质(包括胺、氨基酸、氨基糖和硝基甲烷分子)可以作为消毒副产物前驱体,HCLL-S8-M滤柱出水中N-DBPs FP低于椰壳活性炭滤柱出水中的含量[28]。由此证明生物滤池处理过程中,微生物群落通过EPS反映环境胁迫[29]。同时证明,HCLL-S8-M滤柱相比椰壳活性炭滤柱可以削弱复合微污染物对微生物的胁迫效应,微生物不需要过度分泌EPS来抵御环境胁迫,减弱微生物之间的抗性基因水平转移[30]。
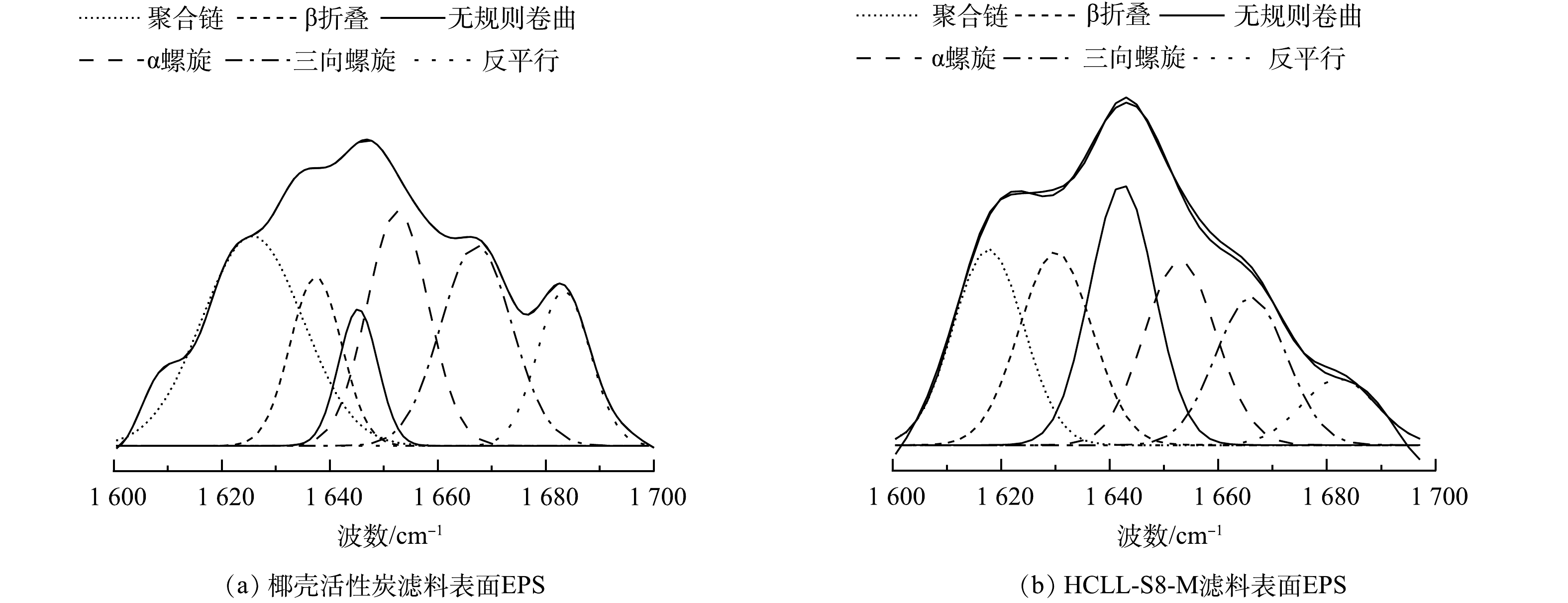
通过拟合红外光谱酰胺I区,得到滤料表面EPS中蛋白质二级结构(图7,表1)。通常α-螺旋和β-折叠占比越大,EPS的疏水性就越强,同时蛋白质与多糖的比值可以判断疏水性的强弱。 HCLL-S8-M滤柱中EPS样品中的α-螺旋和β-折叠占比之和为37.23%,高于椰壳活性炭滤柱的32.06%,同时与图5(a)中PN/PS一致,表明HCLL-S8-M滤柱中EPS疏水性更强[21]。因此,HCLL-S8-M滤柱中EPS具有高凝聚效率能够稳定附着在滤料表面,减小其脱落水体中风险从而控制消毒副产物前驱体的生成。
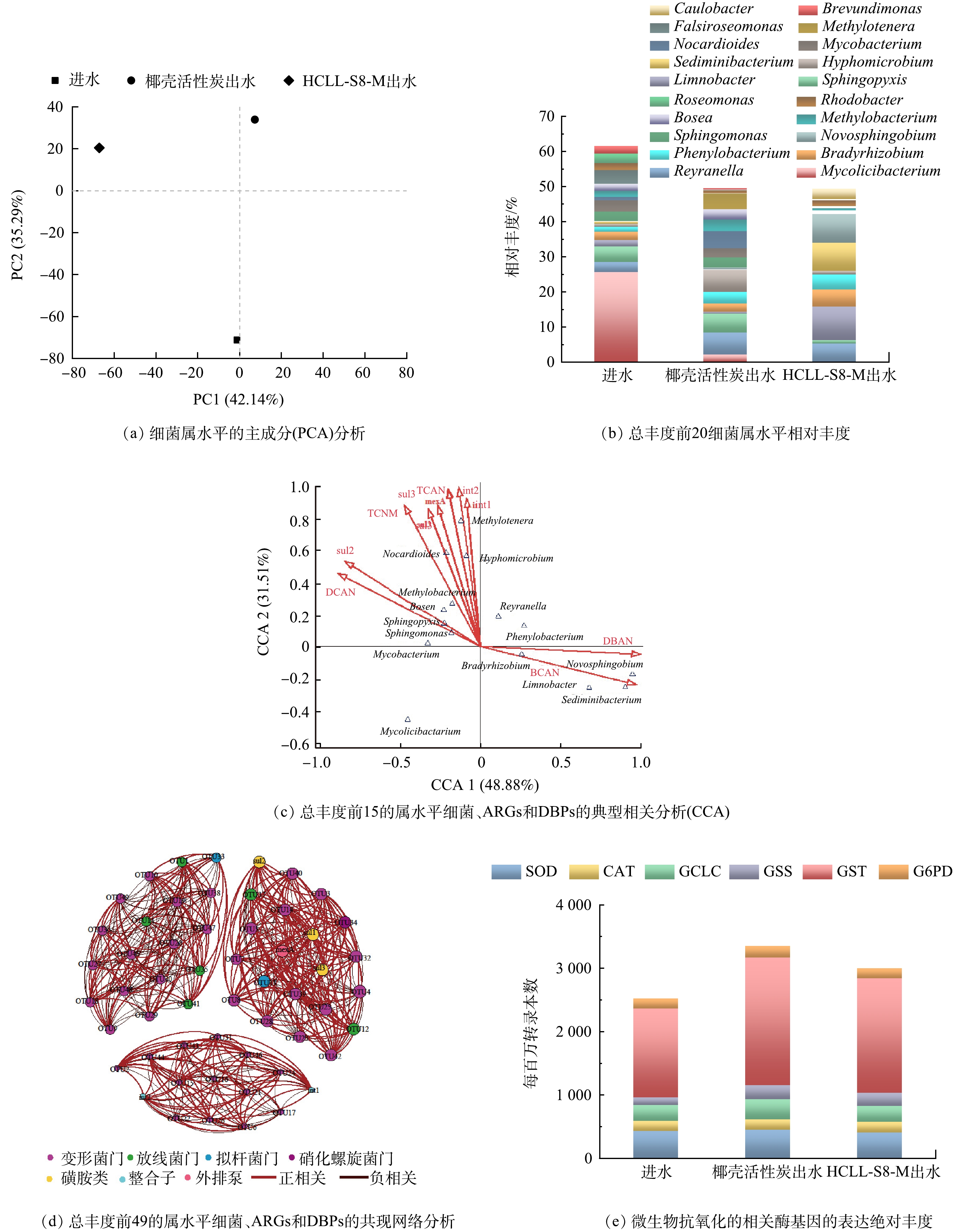
宏基因组学分析细菌属水平的PCA(图8(a))证实滤料对微生物群落结构有显著影响。图8(b)证明进水中分枝菌酸杆形菌属(Mycolicibacterium)占比最大(25.60%),滤料处理后出水中占比小于3%。HCLL-S8-M滤柱出水中,湖沉积杆菌(Limnobacter)、沉积物杆状菌属(Sediminibacterium)和新鞘酯属(Novosphingobium)成为优势细菌属。另外生丝微菌属(Hyphomicrobium)、鞘氨醇盒菌属(Sphingopyxis)、类诺卡氏菌属(Nocardioides)和Methylotenera在椰壳活性炭滤柱出水微生物中占比分别为6.27%、5.35%、4.89%和4.49成为优势菌群。通过CCA分析(图8(c))可以得出HCLL-S8-M滤料出水中的优势细菌属只与DBAN和BCAN成正相关,并且这2种N-DBPs在检测出的消毒副产物中占比最小。椰壳活性炭滤料出水中的优势菌群与ARGs和N-DBPs均有显著的相关性。共现网络分析结果(图8(d))用于说明ARGs与总丰度前49的菌群(表2)之间的关系。13个OTU(例如OTU为3、7、13、14、25)与磺胺类抗性基因和外排泵呈正相关,OTU2、OTU 21等10个OTU与整合子显著相关,表明这些OTU是抗性基因的潜在宿主。结合表2细菌群落绝对丰度发现,抗性基因的潜在宿主的绝对丰度在HCLL-S8-M滤柱出水微生物中要远低于椰壳活性炭滤柱出水微生物。这表明HCLL-S8-M滤料通过改变微生物群落削弱抗性基因的生成和水平转移。微生物受到环境胁迫时,会引发氧化应激反应,维持正常存活率。超氧化物歧化酶(SOD)、过氧化氢酶(CAT)、谷氨酸-半胱氨酸连接酶(GCLC)、谷胱甘肽合成酶(GSS)、谷胱甘肽S转移酶(GST)、葡萄糖-6-磷酸脱氢酶(G6PD)和谷胱甘肽过氧化物酶(GPX)参与氧化应激反应调节细胞内活性氧(ROS)。图8(e)显示HCLL-S8-M滤柱出水中微生物抗氧化应激反应相关酶的总体绝对丰度低于椰壳活性炭滤柱出水中。在相同的复合微污染物条件下,水体中复杂的有机物可能会诱发微生物氧化应激反应,破坏抗氧化系统的平衡,诱发氧化损伤,促进EPS分泌[31]。在HCLL-S8-M滤柱出水中微生物功能基因下调,表明HCLL-S8-M滤料可以削弱微污染物对微生物带来的胁迫效应,降低微生物氧化应激和EPS分泌,从而减少DBPs前驱体并抑制ARGs的水平转移[32-33]。
-
1) HCLL-S8-M滤料去除N-DBPs前驱体和ARGs的效率远高于椰壳活性炭滤料,出水中N-DBPs FP的总量比椰壳活性炭滤柱出水低25.74%,同时出水中ARGs的相对丰度(4.77%)比椰壳活性炭滤柱出水(17.27%)更低。此外,HCLL-S8-M滤料可以有效去除包括微污染物在内的DOC,起到净化饮用水水质作用。
2)在HCLL-S8-M滤柱中微生物分泌更少的EPS,并且其能够更加稳定附着在滤料表面。HCLL-S8-M滤柱通过重组微生物群落,降低ARGs潜在宿主的数量,同时微生物氧化应激反应功能基因表达下降,表明HCLL-S8-M滤柱可以削弱微污染物对微生物带来的胁迫效应,削弱微生物分泌EPS,因此,DBPs前驱体降低并抑制ARGs的水平转移。
催化剂填料表面微生物对微污染物胁迫的响应
Response of catalyst microorganisms to the stress effects of micropollutants
-
摘要: 目前复合微污染水源条件下,饮用水中复合微污染物对微生物带来的胁迫效应会引发含氮类消毒副产物生成势(nitrogenous disinfection byproducts formation potential, N-DBPs FP)和抗性基因(antibiotic resistance genes, ARGs)水质风险。本研究基于催化剂(HCLL-S8-M)作为新型生物滤料建立反应器处理砂滤后水,考察生物滤池出水N-DBPs FP和ARGs丰度,对滤料胞外聚合物(extracellular polymeric substance, EPS)进行分析,通过宏基因组学分析水体中微生物群落结构和胁迫效应基因表达水平。结果表明,在相同复合微污染物条件下(磺胺嘧啶和2,4-二氯苯氧乙酸各1 µg·L−1),HCLL-S8-M滤柱出水中卤代乙腈和卤代硝基甲烷的总生成势为522.30 ng·L−1,比椰壳活性炭滤柱出水低171.70 ng·L−1,ARGs相对丰度也比椰壳活性炭滤柱出水低约4倍。这得益于HCLL-S8-M滤柱可以高效去除包括微污染物在内的溶解性有机碳、SUVA和高分子质量的有机物。HCLL-S8-M滤柱和出水中微生物分泌更少的EPS,从而减少的N-DBPs前驱体和削弱ARGs的水平转移。宏基因组学分析结果表明,微生物在HCLL-S8-M滤柱出水中的氧化应激反应功能基因表达比椰壳活性炭滤柱下降。HCLL-S8-M滤柱减弱微污染物对微生物的胁迫效应,降低N-DBPs FP和ARGs,有助于控制微生物应激反应造成间接饮用水水质风险。Abstract: Under the current conditions of composite micro-polluted water sources, the stress effects of composite micropollutants in drinking water for microbes can cause water quality risks of nitrogenous disinfection byproducts formation potential (N-DBPs FP) and antibiotic resistance genes (ARGs). In this study, the catalyst HCLL-S8-M was used as a new biofilter to build the reactor treating the effluent from sand filtration. The abundances of N-DBPs FP and ARGs in the biofilter effluent were studied, the filter media and extracellular polymeric substance (EPS) was analyzed, and metagenome sequencing was used to analyze the microbial community structure and gene expression levels of stress effect in water bodies. The results showed that under the exposure of the same complex micropollutants conditions (1 µg·L−1 of sulfadiazine and 1 µg·L−1 of 2,4-dichlorophenoxyacetic acid), the total formation potential of haloacetonitriles and halonitromethanes was 522.30 ng·L−1, which was 171.70 ng·L−1 lower than that of coconut shell activated carbon, the relative abundance of ARGs was around 4 times lower than that of coconut shell activated carbon effluent. This was due to HCLL-S8-M could efficiently remove dissolved organic carbon (including micropollutants), SUVA and organics with high molecular weight. Microorganisms in the HCLL-S8-M filter and effluent secreted few EPS, which resulted in reduced precursors of N-DBPs and horizontal transfer of ARGs. Metagenome analysis revealed that oxidative stress functional gene expression of microorganisms decreased on HCLL-S8-M compared to coconut shell activated carbon. HCLL-S8-M attenuated the stress effects of micropollutants on microorganisms, and decreased N-DBPs FP and ARGs, which will contribute to control the indirect drinking water quality risks of micropollutants from the microbial stress response during biofiltration.
-
-
图 8 微生物属水平的PCA分析、前20种细菌的相对丰度、典型相关分析、共现网络分析和抗氧化相关酶基因绝对丰度
Figure 8. PCA analysis of microbes at genus level, relative abundances of the top 20 bacteria, canonical correlation analysis, co-occurrence network analysis and expression profiles for related enzyme genes of antioxidant with absolute abundance
表 1 滤料表面EPS蛋白质二级结构占比情况
Table 1. Percentage of protein secondary structures of EPS on filters surface
蛋白质二级结构 波数/cm−1 蛋白质二级结构所占比例% 椰壳活性炭 HCll-S8-M 聚合链 1 625~1 610 29.30 18.79 β-折叠 1 640~1 630 12.07 19.13 无规则卷曲 1 645~1 640 7.44 22.90 α-螺旋 1 657~1 648 19.99 18.10 三向螺旋 1 672~1 659 19.80 14.36 反平行/β-折叠 1 695~1 680 11.42 6.71 表 2 图8(d)中OTU对应的门、属和绝对丰度
Table 2. Phylum, Genus and absolute abundance corresponding to OTU in Fig.8(d)
OTU 门 属 进水 椰壳活性炭滤柱出水 HCLL-S8-M滤柱出水 OTU1 Actinobacteria Mycolicibacterium 11256 836 813 710 48 896 OTU2 Proteobacteria Reyranella 1293 900 2435 894 1453 014 OTU3 Proteobacteria Sphingopyxis 1910 720 2058 776 266 966 OTU4 Proteobacteria Limnobacter 811 708 207 092 2695 466 OTU5 Proteobacteria Bradyrhizobium 1039 726 901 014 1369 452 OTU6 Proteobacteria Phenylobacterium 622 486 1255 288 1228 110 OTU7 Proteobacteria Hyphomicrobium 396 548 2410 284 273 686 OTU8 Bacteroidota Sediminibacterium 253 564 70 144 2271 874 OTU9 Proteobacteria Novosphingobium 66 532 213 954 2314 156 OTU10 Proteobacteria Sphingomonas 1183 450 1111 608 259 714 OTU11 Actinobacteria Mycobacterium 1410 282 989 004 32 716 OTU12 Actinobacteria Nocardioides 420 912 1880 760 5 356 OTU13 Proteobacteria Methylobacterium 787 802 1274 470 170 738 OTU14 Proteobacteria Bosea 824 698 1122 304 97 046 OTU15 Proteobacteria Methylotenera 29 230 1726 124 59 582 OTU16 Proteobacteria Falsiroseomonas 1738 764 12 182 21 782 OTU17 Proteobacteria Rhodobacter 860 160 340 022 446 816 OTU18 Proteobacteria Roseomonas 1223 196 74 974 31 470 OTU19 Proteobacteria Brevundimonas 928 140 178 374 84 584 OTU20 Proteobacteria Caulobacter 69 884 113 844 797 672 OTU21 Proteobacteria Methylophilus 114 028 524 408 333 398 OTU22 Proteobacteria Tabrizicola 504 872 201 308 263 828 OTU23 Proteobacteria Piscinibacter 77 592 61 154 826 228 OTU24 Proteobacteria Aquisediminimonas 1 130 948 778 3 852 OTU25 Proteobacteria Sphingobium 187 780 628 680 137 014 OTU26 Proteobacteria Curvibacter 12 124 695 130 112 822 OTU27 Proteobacteria Afipia 77 162 326 084 409 316 OTU28 Proteobacteria Aestuariivirga 291 710 435 756 81 166 OTU29 Proteobacteria Solimonas 619 750 87 994 17 044 OTU30 Proteobacteria Aquabacterium 15 084 98 008 529 220 OTU31 Proteobacteria Methyloversatilis 3 124 477 578 42 298 OTU32 Proteobacteria Mesorhizobium 156 102 248 982 41 414 OTU33 Bacteroidota Haliscomenobacter 380 072 53 280 1 696 OTU34 Nitrospirae Nitrospira 61 004 298 250 27 688 OTU35 Actinobacteria Pimelobacter 279 370 83 954 550 OTU36 Proteobacteria Methylibium 11 040 83 316 268 244 OTU37 Actinobacteria Miltoncostaea 21 526 331 566 4 632 OTU38 Proteobacteria Rubrivivax 139 984 113 738 96 944 OTU39 Proteobacteria Rhodoplanes 11 736 251 922 9 530 OTU40 Proteobacteria Pseudomonas 80 188 101 058 74 944 OTU41 Actinobacteria Mycobacteroides 128 118 106 338 20 612 OTU42 Proteobacteria Pseudorhodoplanes 45 956 160 012 44 112 OTU43 Proteobacteria Legionella 19 490 194 794 29 928 OTU44 Proteobacteria Rhodoferax 17 382 165 722 48 922 OTU45 Proteobacteria Hydrogenophaga 22 826 70 754 76 062 OTU46 Proteobacteria Methylorubrum 6 972 154 324 8 248 OTU47 Proteobacteria Acidovorax 84 840 38 906 34 702 OTU48 Proteobacteria Ideonella 19 214 26 610 52 772 OTU49 Proteobacteria Aquariibacter 856 2 218 9 060 -
[1] ALSBAIEE A, SMITH B J, XIAO L, et al. Rapid removal of organic micropollutants from water by a porous beta-cyclodextrin polymer[J]. Nature, 2016, 529: 190-194. doi: 10.1038/nature16185 [2] SU Z, YU W, LIU T, et al. Discovery of welcome biopolymers in surface water: Improvements in drinking water production[J]. Environmental Science & Technology, 2021, 55: 2076-2086. [3] JIA S, BIAN K, SHI P, et al. Metagenomic profiling of antibiotic resistance genes and their associations with bacterial community during multiple disinfection regimes in a full-scale drinking water treatment plant[J]. Water Research, 2020, 176: 115721. doi: 10.1016/j.watres.2020.115721 [4] FANG Y, CHEN C, CUI B, et al. Self-rescue of nitrogen-cycling bacteria under beta-lactam antibiotics stress during managed aquifer recharge (MAR): Microbial collaboration and anti-resistance[J]. Water Research, 2023, 231: 119623. doi: 10.1016/j.watres.2023.119623 [5] CAREY D E, MCNAMARA P J. Altered antibiotic tolerance in anaerobic digesters acclimated to triclosan or triclocarban[J]. Chemosphere, 2016, 163: 22-26. doi: 10.1016/j.chemosphere.2016.07.097 [6] GAZE W H, ABDOUSLAM N, HAWKEY P M, et al. Incidence of class 1 integrons in a quaternary ammonium compound-polluted environment[J]. Antimicrob Agents Chemother, 2005, 49: 1802-1807. doi: 10.1128/AAC.49.5.1802-1807.2005 [7] KURENBACH B, MARJOSHI D, AMABILE-CUEVAS C F, et al. Sublethal exposure to commercial formulations of the herbicides dicamba, 2, 4-dichlorophenoxyacetic acid, and glyphosate cause changes in antibiotic susceptibility in Escherichia coli and Salmonella enterica serovar Typhimurium[J]. mBio, 2015, 6: e00009-00015. [8] 董丽华, 陈志颖, 韩英杰, 等. 新疆煤基压块活性炭在BAC工艺中的应用[J]. 环境工程学报, 2023, 17(8): 2587-2595. [9] FU J, LEE W N, COLEMAN C, et al. Removal of disinfection byproduct (DBP) precursors in water by two-stage biofiltration treatment[J]. Water Research, 2017, 123: 224-235. doi: 10.1016/j.watres.2017.06.073 [10] XIN K, CHEN X, ZHANG Z, et al. Trace antibiotics increase the risk of antibiotic resistance genes transmission by regulating the biofilm extracellular polymeric substances and microbial community in the sewer[J]. Journal of Hazardous Materials, 2022, 432: 128634. doi: 10.1016/j.jhazmat.2022.128634 [11] 胡炽盛, 马徐, 倪炯, 等. 微量磺胺甲恶唑对饮用水管网生物膜群落及抗性基因的影响与控制[J]. 环境工程学报, 2023, 17(6): 2077-2087. [12] YAN X, LIN T, WANG X, et al. Effects of pipe materials on the characteristic recognition, disinfection byproduct formation, and toxicity risk of pipe wall biofilms during chlorination in water supply pipelines[J]. Water Research, 2022, 210: 117980. doi: 10.1016/j.watres.2021.117980 [13] 蒋柱武, 刘欣汝, 武江南, 等. 饮用水中典型含氮消毒副产物的生成与控制研究进展[J]. 环境工程学报, 2020, 14(10): 2595-2603. [14] LI J, YE W, WEI D, et al. System performance and microbial community succession in a partial nitrification biofilm reactor in response to salinity stress[J]. Bioresource Technology, 2018, 270: 512-518. doi: 10.1016/j.biortech.2018.09.068 [15] WANG Z, CHOI O, SEO Y. Relative contribution of biomolecules in bacterial extracellular polymeric substances to disinfection byproduct formation[J]. Environmental Science & Technology, 2013, 47: 9764-9773. [16] LEE W, WESTERHOFF P, CROUE J P. Dissolved organic nitrogen as a precursor for chloroform, dichloroacetonitrile, N-nitrosodimethylamine, and trichloronitromethane[J]. Environmental Science & Technology, 2007, 41: 5485-5490. [17] LYU L, YAN D B, YU G F, et al. Efficient destruction of pollutants in water by a dual-reaction-center fenton-like process over carbon nitride compounds-complexed Cu(II)-CuAlo2[J]. Environmental Science & Technology, 2018, 52(7): 4294-4304. [18] LYU L, YU G F, ZHANG L L, et al. 4-phenoxyphenol-functionalized reduced graphene oxide nanosheets: A metal-free fenton-like catalyst for pollutant destruction[J]. Environmental Science & Technology, 2018, 52(7): 747-756. [19] TSELIOS C, PAPAGEORGIOU M, VAROTSIS C. Extracellular electron uptake from carbon-based pi electron surface-donors: oxidation of graphite sheets by Sulfobacillus thermosulfidooxidans probed by Raman and FTIR spectroscopies[J]. The Royal Society of Chemistry, 2019, 9(33): 19121-19125. [20] LYU L, ZHANG G, HE H, et al. Selective H2O2 conversion to hydroxyl radicals in the electron-rich area of hydroxylated C-g-C3N4/CuCo-Al2O3[J]. Journal of Materials Chemistry, 2017, 5: 7153-7164. doi: 10.1039/C7TA01583F [21] GAO J Y, XING X C, CAI W, et al. Effect of micropollutants on disinfection byproducts and antibiotic resistance genes in drinking water in the process of biological activated carbon treatment[J]. Journal of Hazardous Materials, 2024, 461: 132304. doi: 10.1016/j.jhazmat.2023.132304 [22] ZHANG R, WANG F, CHU W, et al. Microbial degradation of typical amino acids and its impact on the formation of trihalomethanes, haloacetonitriles and haloacetamides during chlor(am)ination[J]. Water Research, 2019, 159: 55-64. doi: 10.1016/j.watres.2019.04.032 [23] WANG H B, HU C, LIU L Z, et al. Interaction of ciprofloxacin chlorination products with bacteria in drinking water distribution systems[J]. Journal of Hazardous Materials, 2017, 339: 174-181. doi: 10.1016/j.jhazmat.2017.06.033 [24] ZHENG J, LIN T, CHEN W, et al. Removal of precursors of typical nitrogenous disinfection byproducts in ozonation integrated with biological activated carbon (O3/BAC)[J]. Chemosphere, 2018, 209: 68-77. doi: 10.1016/j.chemosphere.2018.06.018 [25] JIA F, YANG Q, LIU X, et al. Stratification of Extracellular Polymeric Substances (EPS) for Aggregated Anammox Microorganisms[J]. Environmental Science & Technology, 2017, 51: 3260-3268. [26] HUA G, RECKHOW D A, ABUSALLOUT I, et al. Correlation between SUVA and DBP formation during chlorination and chloramination of NOM fractions from different sources[J]. Chemosphere, 2015, 130: 82-89. doi: 10.1016/j.chemosphere.2015.03.039 [27] CHUNG Y, KIM H, KIM T S, et al. Mitigation of organic fouling on ceramic membranes by selective removal of microbial-oriented organic matters in wastewater effluents[J]. Separation and Purification Technology, 2019, 219: 216-221. doi: 10.1016/j.seppur.2019.03.032 [28] ZHENG J, CHEN T, CHEN H. Antibiotic resistome promotion in drinking water during biological activated carbon treatment: Is it influenced by quorum sensing?[J]. Environmental Science & Technology, 2018, 612: 1-8. [29] SHAH A D, MITCH W A. Halonitroalkanes, halonitriles, haloamides, and N-nitrosamines: a critical review of nitrogenous disinfection byproduct formation pathways[J]. Environmental Science & Technology, 2012, 46: 119-131. [30] MA W J, REN Z Q, YU L Q, et al. Deciphering the response of anammox process to heavy metal and antibiotic stress: Arsenic enhances the permeability of extracellular polymeric substance and aggravates the inhibition of sulfamethoxazole[J]. Chemical Engineering Journal, 2021, 426: 130815. doi: 10.1016/j.cej.2021.130815 [31] XING X C, LYU L, YAN Z, et al. Self-purification of actual wastewater via microbial-synergy driving of catalyst-surface microelectronic field: A pilot-scale study[J]. Journal of Hazardous Materials, 2023, 457: 131744. doi: 10.1016/j.jhazmat.2023.131744 [32] WANG Y H, WU Y H, LUO L W, et al. Metagenomics analysis of the key functional genes related to biofouling aggravation of reverse osmosis membranes after chlorine disinfection[J]. Journal of Hazardous Materials, 2021, 410: 124602. doi: 10.1016/j.jhazmat.2020.124602 [33] DI TOMMASO C, TAYLOR-EDMONDS L, ANDREWS S A, et al. The contribution of biofilm to nitrogenous disinfection by-product formation in full-scale cyclically-operated drinking water biofilters[J]. Water Research, 2019, 155: 403-409. doi: 10.1016/j.watres.2019.02.025 -


 下载:
下载: